The follow is a guest post by Bob Mesibov, who has contributed to iPhylo before. 
Like many iPhylo readers, I deal with large, pre-existing compilations of biodiversity data. The compilations come from museums, herbaria, aggregation projects and government agencies. For simplicity in what follows and to avoid naming names, I'll lump all these sources into a single fictional entity, the PAI (for Projects, Agencies and Institutions).
The datasets I get from the PAI typically contain duplicate records, inconsistencies in content and format, unexplained data gaps, data in wrong fields, fields improperly used, no flagging of doubtful data, etc. Data cleaning consumes most of the time I spend on a data project. Cleaning can take weeks, analysing the cleaned data takes minutes, reporting the results of the analysis takes hours or days. (Example: doi:10.3897/BDJ.2.e1160)
I can understand how datasets get messy. Data entry errors account for a lot of the problems I see, and I make data entry errors myself. But the causes of messiness are one thing and its cure is another. The custodians of those data compilations don't seem to have done much (or any) data checking. Why not?
When I'm brave enough to ask that question, I usually get a polite response from the PAI. Here are 10 explanations I've heard for inadequate data checking and cleaning:
(1) The data are fit for use, as-is.
No cleaning is needed, because the data are fit for some use, and the PAI is satisfied with that. One data manager wrote to me in an email: '...even records with lower certainty, in this case an uncertain identification, can be useful at a coarser resolution. Although we have no idea as to the reliability of the identification to the species or even genus they are likely correctly identify[ing] something as at least an animal, arthropod and possibly to class so the record is suitable for analysis at that level.'
(2) The PAI is exposing its data online. The crowd will spot any problems and tell the PAI about them.
I've previously pointed out (doiL10.3897/zookeys.293.5111) how lame this explanation is. As a strategy for data cleaning it's slow, piecemeal and wildly optimistic. At best, it accumulates data-cleaning 'tickets' with no guarantee that any will ever be closed. What I hear from the PAI is 'We're aware of problems of that kind and are hoping to find a general solution, rather than deal with a multitude of individual cases'. Years pass and the individual cases don't get fixed, so interested members of the crowd lose faith in the process and stop reporting problems.
(3) No one outside the PAI is allowed to look at the whole dataset, and no one inside the PAI has the time (or skills) to do data checking and cleaning.
This is a particularly nice Catch-22. I once offered to check a portion of the PAI's data holdings for free, and was told that PAI policy was that the dataset was not to be shared with anyone outside the PAI. The same data were freely available on the PAI's website in bits and pieces through a database search page.
(4) The PAI is migrating to new database software next year. Data cleaning will be part of the migration.
No, it won't. Note that this response isn't always simple procrastination, because it's sometimes the case that the PAI's database has only limited capabilities for data checking and editing. PAI staff are hopeful that checking and editing will be easier with the new software. They'll be disappointed.
(5) The person who manages data is on leave / was seconded to another project / resigned and hasn't been replaced yet / etc.
This is another way of saying that no one inside the PAI has the time to do data checking and cleaning. When the data manager returns to work or gets replaced, data checking and cleaning will have the same low priority it had before. That's why it didn't get done.
(6) Top management says any data cleaning would have to be done by outside specialists, but there's not enough money in the current budget to hire such people.
Not only a Catch-22, but a solid, long-term excuse, applicable in any financial year. It would cost less to train PAI staff to do the job in-house.
(7) The PAI would prefer to use a specialist data tool to clean data, like OpenRefine, but hasn't yet got up to speed on its use.
The PAI believes in magic. OpenRefine will magically clean the data without any thought required on the part of PAI staff. The magic will have to be applied repeatedly, because the sources of the duplications, gaps and errors haven't been found and squashed.
(8) The PAI staff best qualified to check and clean the data aren't allowed to do so.
IT policy strictly forbids anyone but IT staff from tinkering with the PAI database, whose integrity is sacrosanct. A very specific request from biodiversity staff may be ticketed by IT staff for action, but global checking and editing is out of the question. IT staff are not expected to understand biodiversity studies, and biodiversity staff are not expected to understand databases.
This explanation is interesting because it implies a workaround. If a biodiversity staffer can get a dump from the database as a simple text file, she can do global checking and editing of the data using the command line or a spreadsheet. The cleaned data can then be passed to IT staff for incorporation into the database as replacement data items. The day that happens, pigs will be seen flying outside the PAI windows.
(9) The PAI datasets have grown so big that global data checking and editing is no longer possible.
Harder, yes; impossible, no. And the datasets didn't suddenly appear, they grew by accretion. Why wasn't data checking and editing done as data was added?
(10) All datasets are messy and data users should do their own data cleaning.
The PAI shrugs its shoulders and says 'That's just the way it is, live with it. Our data are no messier than anyone else's'.
I've left this explanation for last because it begs the question. Yes, users can do their own data cleaning — because it's not that hard and there are many ways to do it. So why isn't it done by highly qualified, well-paid PAI data managers?
 Despite the well deserved scepticism about dashboards voiced by Shannon Mattern @shannonmattern (see Mission Control: A History of the Urban Dashboard, I discovered this by reading Ignore the Bat Caves and Marketplaces: lets talk about Zoning by Leigh Dodds @ldodds) I'm intrigued by the idea a dashboard for biodiversity. We could have several different kinds of information, displayed in a single place.
Despite the well deserved scepticism about dashboards voiced by Shannon Mattern @shannonmattern (see Mission Control: A History of the Urban Dashboard, I discovered this by reading Ignore the Bat Caves and Marketplaces: lets talk about Zoning by Leigh Dodds @ldodds) I'm intrigued by the idea a dashboard for biodiversity. We could have several different kinds of information, displayed in a single place. For anyone doing research or involved in scientific infrastructure, demonstrating the "impact" of those activities is becoming increasingly important. This has fostered a growth industry in "alt metrics", tools to track how research gathers attention outside academia (of course, we can argue whether attention is the same as impact).
For anyone doing research or involved in scientific infrastructure, demonstrating the "impact" of those activities is becoming increasingly important. This has fostered a growth industry in "alt metrics", tools to track how research gathers attention outside academia (of course, we can argue whether attention is the same as impact).
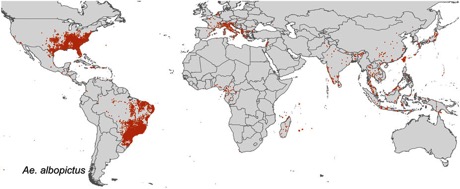

 After experimenting with a dynamic, online version of my notes "Towards a biodiversity knowledge graph" I've published a static version in RIO:
After experimenting with a dynamic, online version of my notes "Towards a biodiversity knowledge graph" I've published a static version in RIO: